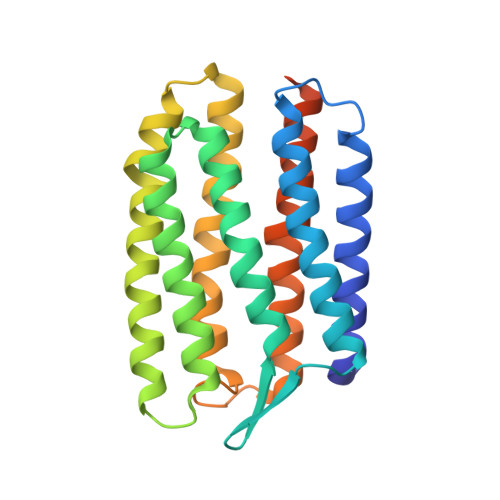
X-Ray Structure of Sensory Rhodopsin II at 2.1 A Resolution
Royant, A., Nollert, P., Edman, K., Neutze, R., Landau, E.M., Pebay-Peyroula, E., Navarro, J.(2001) Proc Natl Acad Sci U S A 98: 10131
- PubMed: 11504917
- DOI: https://doi.org/10.1073/pnas.181203898
- Primary Citation of Related Structures:
1H68 - PubMed Abstract:
Sensory rhodopsins (SRs) belong to a subfamily of heptahelical transmembrane proteins containing a retinal chromophore. These photoreceptors mediate the cascade of vision in animal eyes and phototaxis in archaebacteria and unicellular flagellated algae. Signal transduction by these photoreceptors occurs by means of transducer proteins. The two archaebacterial sensory rhodopsins SRI and SRII are coupled to the membrane-bound HtrI and HtrII transducer proteins. Activation of these proteins initiates phosphorylation cascades that modulate the flagellar motors, resulting in either attractant (SRI) or repellent (SRII) phototaxis. In addition, transducer-free SRI and SRII were shown to operate as proton pumps, analogous to bacteriorhodopsin. Here, we present the x-ray structure of SRII from Natronobacterium pharaonis (pSRII) at 2.1-A resolution, revealing a unique molecular architecture of the retinal-binding pocket. In particular, the structure of pSRII exhibits a largely unbent conformation of the retinal (as compared with bacteriorhodopsin and halorhodopsin), a hydroxyl group of Thr-204 in the vicinity of the Schiff base, and an outward orientation of the guanidinium group of Arg-72. Furthermore, the structure reveals a putative chloride ion that is coupled to the Schiff base by means of a hydrogen-bond network and a unique, positively charged surface patch for a probable interaction with HtrII. The high-resolution structure of pSRII provides a structural basis to elucidate the mechanisms of phototransduction and color tuning.
Organizational Affiliation:
Institut de Biologie Structurale, Unité Mixte de Recherche 5075 Commissariat à l'Energie Atomique-Centre National de la Recherche Scientifique, Université Joseph Fourier, 41 Rue Jules Horowitz, F-38027 Grenoble Cedex 1, France.