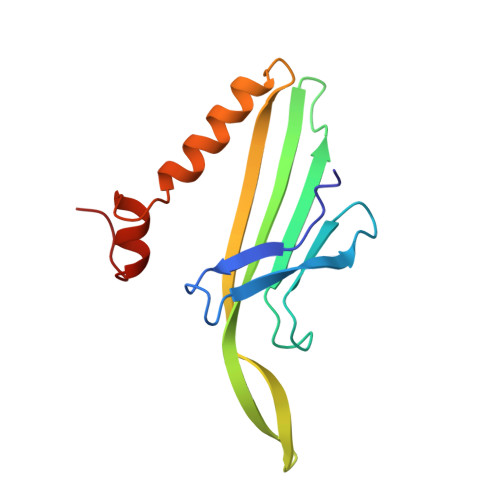
The refined structure of bacteriophage MS2 at 2.8 A resolution.
Golmohammadi, R., Valegard, K., Fridborg, K., Liljas, L.(1993) J Mol Biol 234: 620-639
- PubMed: 8254664
- DOI: https://doi.org/10.1006/jmbi.1993.1616
- Primary Citation of Related Structures:
2MS2 - PubMed Abstract:
Bacteriophage MS2 is an icosahedral virus with 180 copies of a coat protein forming a shell around a single-stranded RNA molecule. The coat protein subunits form a lattice with the triangulation number T = 3. The coat protein has a fold which is different from the fold of all other viral coat proteins so far known. It consists of a five-stranded beta sheet facing the inside of the particle, and a hairpin and two helices on the outside. The crystal structure has been refined at 2.8 A resolution. The final R-factor was 0.189 for reflections with F > 2 sigma, and the root-mean-square deviation from idealized bond lengths and bond angles was 0.015 A and 2.9 degrees, respectively. The three chemically identical conformers A, B and C are largely similar. The B conformer has a unique conformation in one loop, which is involved in 5-fold interactions, while the A and C conformers, which are involved in the quasi-6-fold contacts, are similar throughout the structure. One cis-proline has been identified in the B conformer but the corresponding prolines in A and C are of the trans isomer. This residue is conserved within small RNA coliphages and it is proposed that this isomerization enables a less elongated loop (FG) around the 5-fold axis, thus creating a channel. The extensive dimer contact supports the idea of dimers as initial building blocks. An assembly pathway is proposed where five dimers converge into a pentamer and 12 pentamers are linked together with free dimers creating a complete particle.
Organizational Affiliation:
Department of Molecular Biology, Uppsala University Uppsala Biomedical Centre, Sweden.