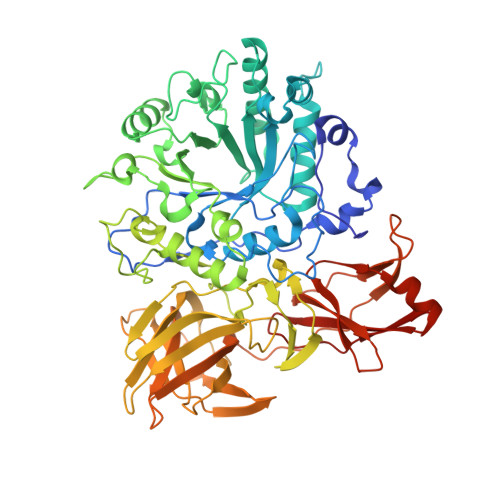
The X-Ray Crystal Structure of an Arthrobacter Protophormiae Endo-Beta-N-Acetylglucosaminidase Reveals a (Beta/Alpha)(8) Catalytic Domain, Two Ancillary Domains and Active Site Residues Key for Transglycosylation Activity.
Suits, M.D., Ling, Z., Bingham, R.J., Bruce, N.C., Davies, G.J., Fairbanks, A.J., Moir, J.W., Taylor, E.J.(2009) J Mol Biol 389: 1
- PubMed: 19327363
- DOI: https://doi.org/10.1016/j.jmb.2009.03.050
- Primary Citation of Related Structures:
2VTF - PubMed Abstract:
Glycoside hydrolase family GH85 is a family of endo-beta-N-acetylglucosaminidases that is responsible for the hydrolysis of beta-1,4 linkage in the N,N-diacetylchitobiose core of N-linked glycans. The endo-beta-N-acetylglucosaminidase from Arthrobacter protophormiae (Endo-A) is of particular interest, given its increasing use for the chemoenzymatic synthesis of bespoke N-glycans using N-glycan oxazolines as glycosyl donors. The E173Q variant of Endo-A is especially attractive for synthesis, as it is hydrolytically impaired but still able to catalyze N-glycan synthesis by transglycosylation using activated oxazoline donors. Here we present the three-dimensional structure of the A. protophormiae Endo-A E173Q variant, solved by multiple-wavelength anomalous scattering methods and refined at 1.8 A resolution. The structure reveals that GH85 enzymes display a trimodular architecture in which a (beta/alpha)(8) catalytic domain occurs with two ancillary beta-sheet modules. The active centre is fully consistent with the known neighboring-group catalytic mechanism in which E173 acts as the catalytic acid/base for reaction via an oxazoline intermediate. Of note is the presence of an asparagine in the active centre, in a position likely to interact with the acetyl NH group that, in all other known families of glycosidase using this mechanism, is an aspartate or glutamate residue. The substrate-binding surface reveals an open topography, consistent with the ability to accept a large range of glycoprotein substrates and the ability to transglycosylate other acceptors. The three-dimensional structure of this important biocatalyst reveals that residues implicated in the enhancement of transglycosylation and synthetic capacity are proximal to the active centre, where they may act to favor binding of acceptor substrates.
Organizational Affiliation:
Department of Biology, University of York, UK.