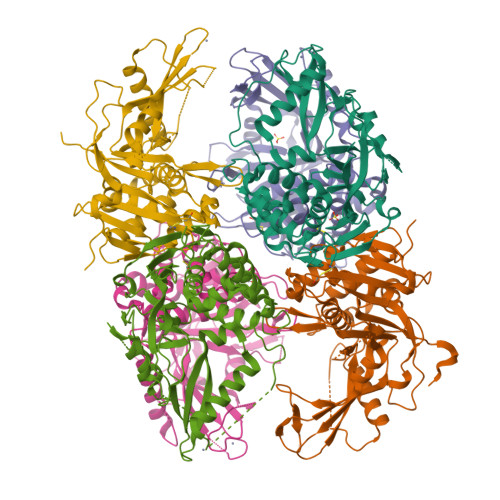
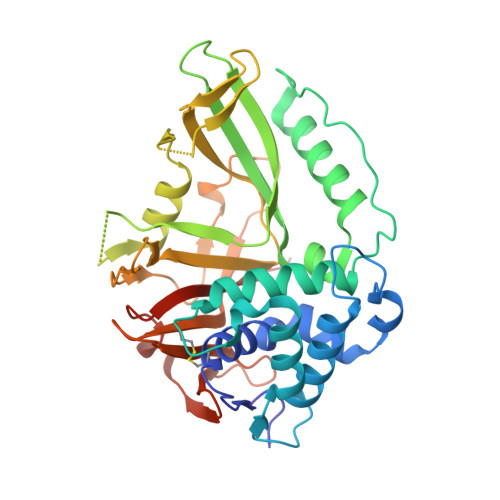
The Dusp-Ubl Domain of Usp4 Enhances its Catalytic Efficiency by Promoting Ubiquitin Exchange.
Clerici, M., Luna-Vargas, M.P.A., Faesen, A.C., Sixma, T.K.(2014) Nat Commun 5: 5399
- PubMed: 25404403
- DOI: https://doi.org/10.1038/ncomms6399
- Primary Citation of Related Structures:
2Y6E - PubMed Abstract:
Ubiquitin-specific protease USP4 is emerging as an important regulator of cellular pathways, including the TGF-β response, NF-κB signalling and splicing, with possible roles in cancer. Here we show that USP4 has its catalytic triad arranged in a productive conformation. Nevertheless, it requires its N-terminal DUSP-Ubl domain to achieve full catalytic turnover. Pre-steady-state kinetics measurements reveal that USP4 catalytic domain activity is strongly inhibited by slow dissociation of ubiquitin after substrate hydrolysis. The DUSP-Ubl domain is able to enhance ubiquitin dissociation, hence promoting efficient turnover. In a mechanism that requires all USP4 domains, binding of the DUSP-Ubl domain promotes a change of a switching loop near the active site. This 'allosteric regulation of product discharge' provides a novel way of regulating deubiquitinating enzymes that may have relevance for other enzyme classes.
Organizational Affiliation:
Department of Biochemistry, The Netherlands Cancer Institute, Plesmanlaan 121, 1066CX Amsterdam, The Netherlands.