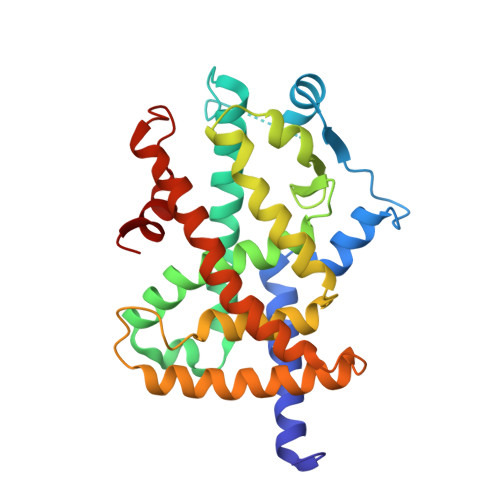
Structural and biochemical basis for the binding selectivity of peroxisome proliferator-activated receptor gamma to PGC-1alpha.
Li, Y., Kovach, A., Suino-Powell, K., Martynowski, D., Xu, H.E.(2008) J Biol Chem 283: 19132-19139
- PubMed: 18469005
- DOI: https://doi.org/10.1074/jbc.M802040200
- Primary Citation of Related Structures:
3CS8 - PubMed Abstract:
The functional interaction between the peroxisome proliferator-activated receptor gamma (PPARgamma) and its coactivator PGC-1alpha is crucial for the normal physiology of PPARgamma and its pharmacological response to antidiabetic treatment with rosiglitazone. Here we report the crystal structure of the PPARgamma ligand-binding domain bound to rosiglitazone and to a large PGC-1alpha fragment that contains two LXXLL-related motifs. The structure reveals critical contacts mediated through the first LXXLL motif of PGC-1alpha and the PPARgamma coactivator binding site. Through a combination of biochemical and structural studies, we demonstrate that the first LXXLL motif is the most potent among all nuclear receptor coactivator motifs tested, and only this motif of the two LXXLL-related motifs in PGC-1alpha is capable of binding to PPARgamma. Our studies reveal that the strong interaction of PGC-1alpha and PPARgamma is mediated through both hydrophobic and specific polar interactions. Mutations within the context of the full-length PGC-1alpha indicate that the first PGC-1alpha motif is necessary and sufficient for PGC-1alpha to coactivate PPARgamma in the presence or absence of rosiglitazone. These results provide a molecular basis for specific recruitment and functional interplay between PPARgamma and PGC-1alpha in glucose homeostasis and adipocyte differentiation.
Organizational Affiliation:
Laboratory of Structural Sciences, Van Andel Research Institute, Grand Rapids, Michigan 49503, USA. yol21@pitt.edu