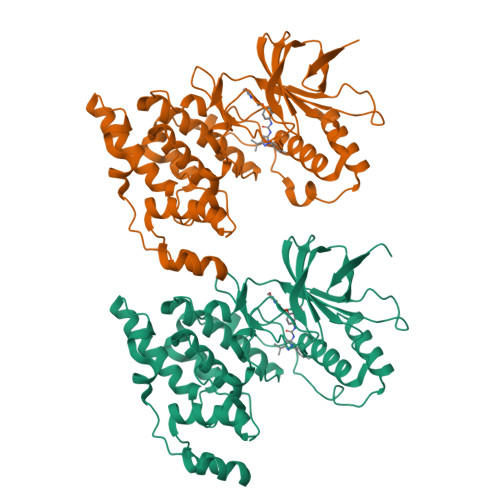
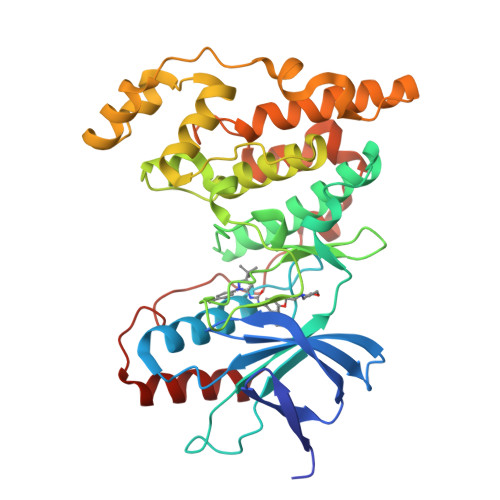
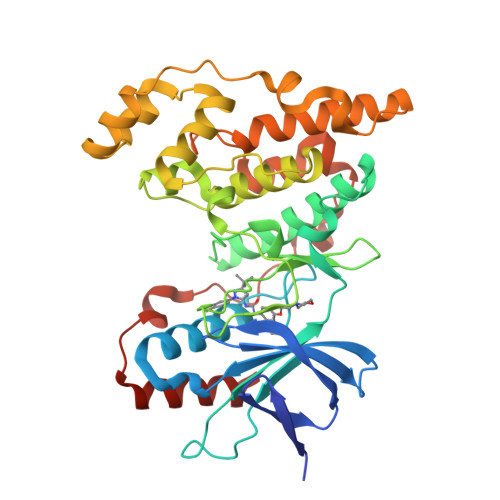
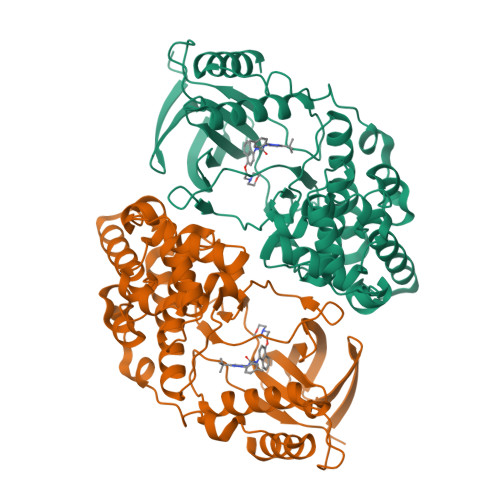
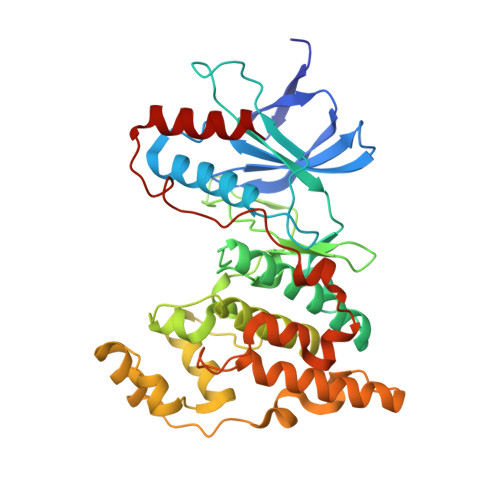
X-ray crystal structure of JNK2 complexed with the p38alpha inhibitor BIRB796: Insights into the rational design of DFG-out binding MAP kinase inhibitors.
Kuglstatter, A., Ghate, M., Tsing, S., Villasenor, A.G., Shaw, D., Barnett, J.W., Browner, M.F.(2010) Bioorg Med Chem Lett 20: 5217-5220
- PubMed: 20655210
- DOI: https://doi.org/10.1016/j.bmcl.2010.06.157
- Primary Citation of Related Structures:
3NPC - PubMed Abstract:
JNK2 and p38alpha are closely related mitogen-activated protein kinases that regulate various cellular activities and are considered drug targets for inflammatory diseases. We have determined the X-ray crystal structure of the clinical phase II p38alpha inhibitor BIRB796 bound to its off-target JNK2. This shows for the first time a JNK subfamily member in the DFG-out conformation. The fully resolved activation loop reveals that BIRB796 inhibits JNK2 activation by stabilizing the loop in a position that does not allow its phosphorylation by upstream kinases. The structure suggests that substituents at the BIRB796 morpholino group and modifications of the t-butyl moiety should further increase the p38alpha to JNK2 potency ratio. For the design of selective DFG-out binding JNK2 inhibitors, the binding pocket of the BIRB796 tolyl group may have the best potential.
Organizational Affiliation:
Roche Palo Alto, Palo Alto, CA 94304, USA. andreas.kuglstatter@roche.com