Discovery, Synthesis, And Structure-Based Optimization of a Series of N-(tert-Butyl)-2-(N-arylamido)-2-(pyridin-3-yl) Acetamides (ML188) as Potent Noncovalent Small Molecule Inhibitors of the Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) 3CL Protease.
Jacobs, J., Grum-Tokars, V., Zhou, Y., Turlington, M., Saldanha, S.A., Chase, P., Eggler, A., Dawson, E.S., Baez-Santos, Y.M., Tomar, S., Mielech, A.M., Baker, S.C., Lindsley, C.W., Hodder, P., Mesecar, A., Stauffer, S.R.(2013) J Med Chem 56: 534-546
- PubMed: 23231439
- DOI: https://doi.org/10.1021/jm301580n
- Primary Citation of Related Structures:
3V3M - PubMed Abstract:
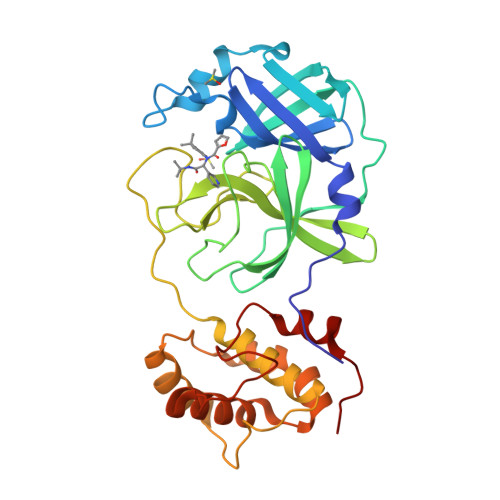
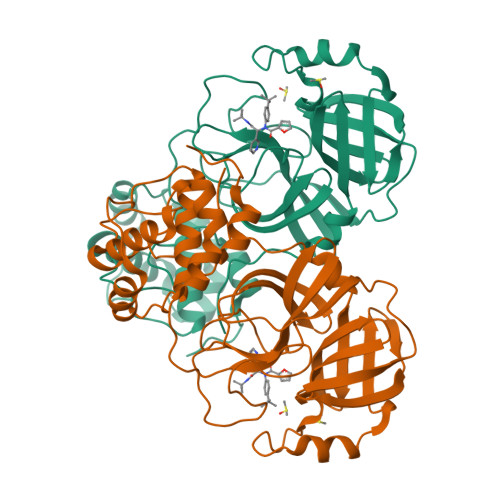
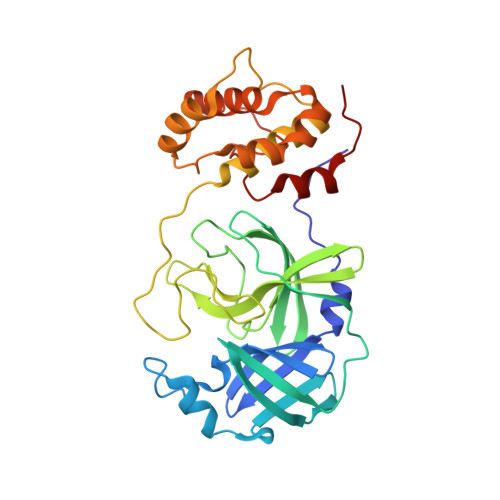
A high-throughput screen of the NIH molecular libraries sample collection and subsequent optimization of a lead dipeptide-like series of severe acute respiratory syndrome (SARS) main protease (3CLpro) inhibitors led to the identification of probe compound ML188 (16-(R), (R)-N-(4-(tert-butyl)phenyl)-N-(2-(tert-butylamino)-2-oxo-1-(pyridin-3-yl)ethyl)furan-2-carboxamide, Pubchem CID: 46897844). Unlike the majority of reported coronavirus 3CLpro inhibitors that act via covalent modification of the enzyme, 16-(R) is a noncovalent SARS-CoV 3CLpro inhibitor with moderate MW and good enzyme and antiviral inhibitory activity. A multicomponent Ugi reaction was utilized to rapidly explore structure-activity relationships within S(1'), S(1), and S(2) enzyme binding pockets. The X-ray structure of SARS-CoV 3CLpro bound with 16-(R) was instrumental in guiding subsequent rounds of chemistry optimization. 16-(R) provides an excellent starting point for the further design and refinement of 3CLpro inhibitors that act by a noncovalent mechanism of action.
Organizational Affiliation:
Department of Pharmacology, Vanderbilt University Medical Center , Nashville, Tennessee 37232, USA.