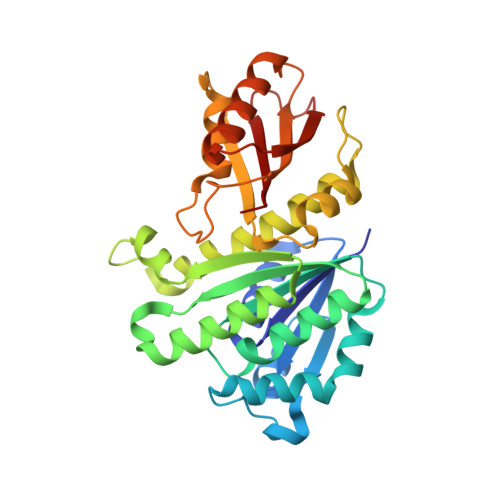
Structural reorganization of the bacterial cell-division protein FtsZ from Staphylococcus aureus
Matsui, T., Yamane, J., Mogi, N., Yamaguchi, H., Takemoto, H., Yao, M., Tanaka, I.(2012) Acta Crystallogr D Biol Crystallogr 68: 1175-1188
- PubMed: 22948918
- DOI: https://doi.org/10.1107/S0907444912022640
- Primary Citation of Related Structures:
3VO8, 3VO9, 3VOA, 3VOB, 3VPA - PubMed Abstract:
FtsZ is a key molecule in bacterial cell division. In the presence of GTP, it polymerizes into tubulin-like protofilaments by head-to-tail association. Protofilaments of FtsZ seem to adopt a straight or a curved conformation in relation to the bound nucleotide. However, although several bacterial and archaeal FtsZ structures have been determined, all of the structures reported previously are considered to have a curved conformation. In this study, structures of FtsZ from Staphylococcus aureus (SaFtsZ) were determined in apo, GDP-bound and inhibitor-complex forms and it was found that SaFtsZ undergoes marked conformational changes. The accumulated evidence suggests that the GDP-bound structure has the features of the straight form. The structural change between the curved and straight forms shows intriguing similarity to the eukaryotic cytoskeletal protein tubulin. Furthermore, the structure of the apo form showed an unexpectedly large conformational change in the core region. FtsZ has also been recognized as a novel target for antibacterial drugs. The structure of the complex with the inhibitor PC190723, which has potent and selective antistaphylococcal activity, indicated that the inhibitor binds at the cleft between the two subdomains.
Organizational Affiliation:
Faculty of Advanced Life Science, Hokkaido University, Sapporo, Hokkaido, Japan.