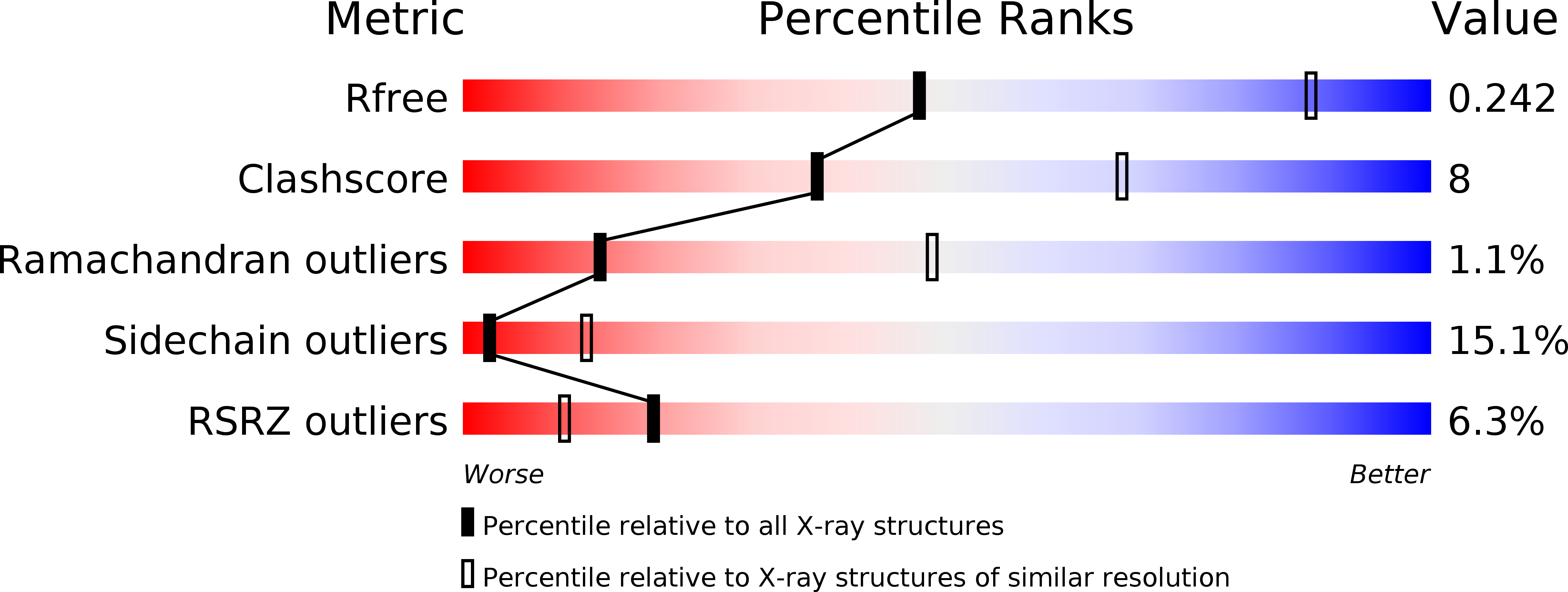
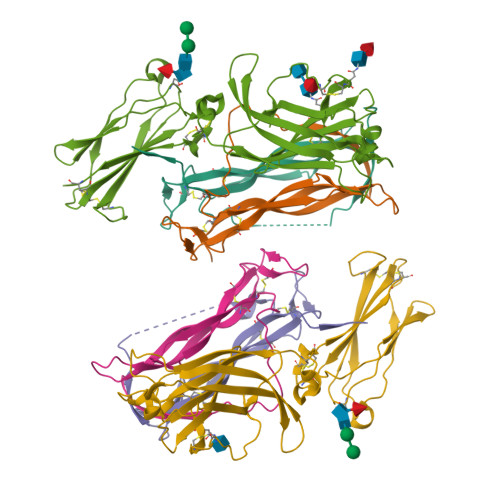
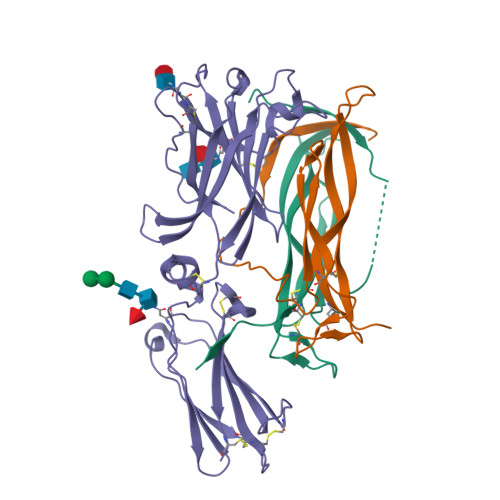
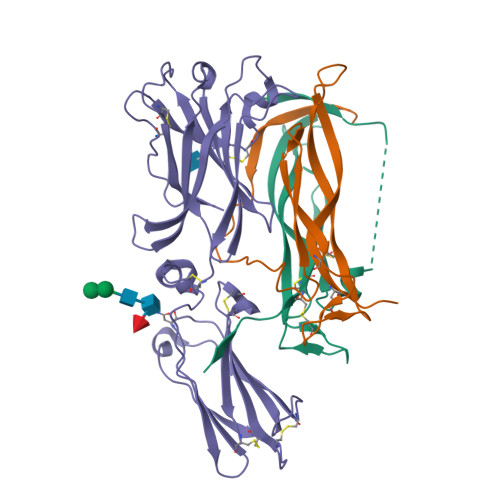
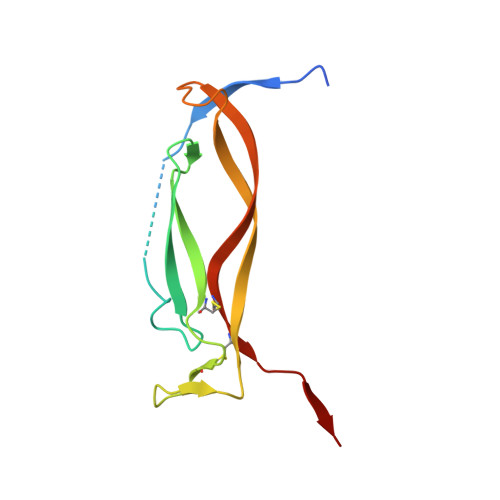
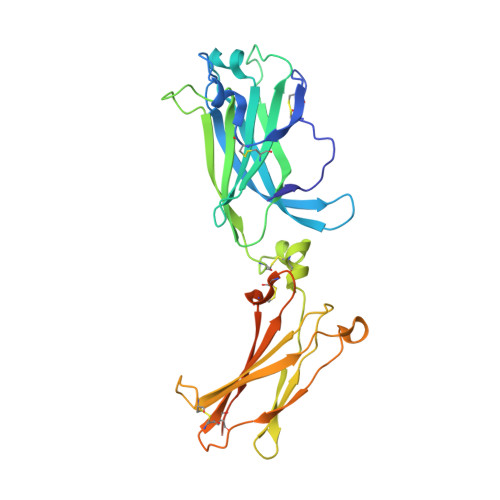
Crystal structures of interleukin 17A and its complex with IL-17 receptor A.
Liu, S., Song, X., Chrunyk, B.A., Shanker, S., Hoth, L.R., Marr, E.S., Griffor, M.C.(2013) Nat Commun 4: 1888-1888
- PubMed: 23695682
- DOI: https://doi.org/10.1038/ncomms2880
- Primary Citation of Related Structures:
4HR9, 4HSA - PubMed Abstract:
The constituent polypeptides of the interleukin-17 family form six different homodimeric cytokines (IL-17A-F) and the heterodimeric IL-17A/F. Their interactions with IL-17 receptors A-E (IL-17RA-E) mediate host defenses while also contributing to inflammatory and autoimmune responses. IL-17A and IL-17F both preferentially engage a receptor complex containing one molecule of IL-17RA and one molecule of IL-17RC. More generally, IL-17RA appears to be a shared receptor that pairs with other members of its family to allow signaling of different IL-17 cytokines. Here we report crystal structures of homodimeric IL-17A and its complex with IL-17RA. Binding to IL-17RA at one side of the IL-17A molecule induces a conformational change in the second, symmetry-related receptor site of IL-17A. This change favors, and is sufficient to account for, the selection of a different receptor polypeptide to complete the cytokine-receptor complex. The structural results are supported by biophysical studies with IL-17A variants produced by site-directed mutagenesis.
Organizational Affiliation:
Structural Biology and Biophysics Group, Pfizer Groton Laboratories, Eastern Point Road, Groton, Connecticut 06340, USA. shenping.liu@pfizer.com