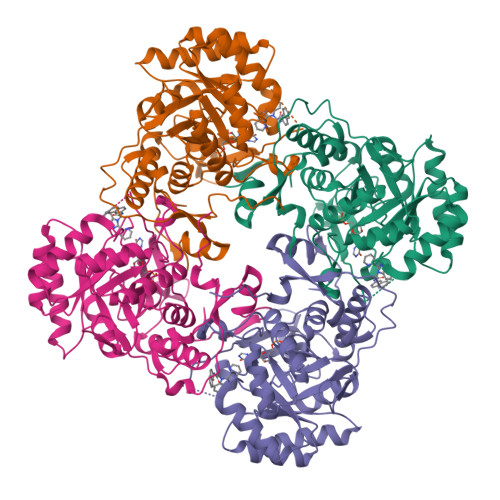
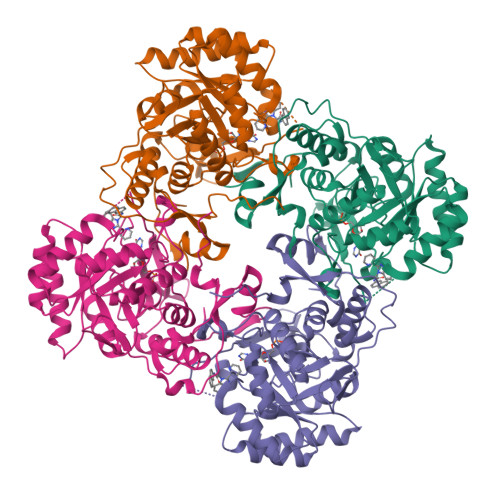
Crystal Structure of Inosine 5'-monophosphate Dehydrogenase from Clostridium perfringens Complexed with IMP and C91
Maltseva, N., Kim, Y., Makowska-Grzyska, M., Mulligan, R., Gu, M., Zhang, M., Mandapati, K., Gollapalli, D.R., Gorla, S.K., Hedstrom, L., Anderson, W.F., Joachimiak, A.To be published.