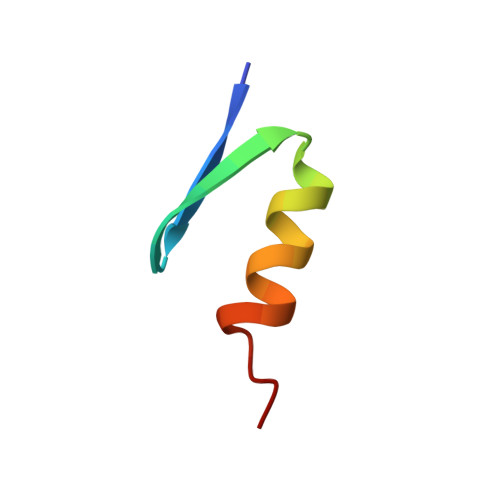
Alternating zinc fingers in the human male associated protein ZFY: 2D NMR structure of an even finger and implications for "jumping-linker" DNA recognition.
Kochoyan, M., Havel, T.F., Nguyen, D.T., Dahl, C.E., Keutmann, H.T., Weiss, M.A.(1991) Biochemistry 30: 3371-3386
- PubMed: 1849423
- DOI: https://doi.org/10.1021/bi00228a004
- Primary Citation of Related Structures:
5ZNF, 7ZNF - PubMed Abstract:
ZFY, a sex-related Zn-finger protein encoded by the human Y chromosome, is distinguished from the general class of Zn-finger proteins by the presence of a two-finger repeat. Whereas odd-numbered domains and linkers fit a general consensus, even-numbered domains and linkers exhibit systematic differences. Because this alternation may have fundamental implications for the mechanism of protein-DNA recognition, we have undertaken biochemical and structural studies of fragments of ZFY. We describe here the solution structure of a representative nonconsensus (even-numbered) Zn finger based on 2D NMR studies of a 30-residue peptide. Structural modeling by distance geometry and simulated annealing (DG/SA) demonstrates that this peptide folds as a miniglobular domain containing a C-terminal beta--hairpin and N-terminal alpha-helix (beta beta alpha motif). These features are similar to (but not identical with) those previously described in consensus-type Zn fingers (derived from ADR1 and Xfin); the similarities suggest that even and odd ZFY domains bind DNA by a common mechanism. A model of the protein-DNA complex (designated the "jumping-linker" model) is presented and discussed in terms of the ZFY two-finger repeat. In this model every other linker is proposed to cross the minor groove by means of a putative finger/linker submotif HX4HX3-hydrophobic residue-X3. Analogous use of a hydrophobic residue in a linker that spans the minor groove has recently been described in crystallographic and 3D NMR studies of homeodomain-DNA complexes. The proposed model of ZFY is supported in part by the hydroxyl radical footprint of the TFIIIA-DNA complex [Churchill, M.E.A., Tullius, T.D., & Klug, A. (1990) Proc. Natl. Acad. Sci. U.S.A. 87, 5528-5532].
Organizational Affiliation:
Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115.